Abstract
Background: In AML and ALL the application of WHO classification and ELN guidelines requires a combination of cytogenetics and targeted sequencing for specific mutations to determine the diagnostic and prognostic subgroup. WGS and WTS have emerged as comprehensive techniques that allow the simultaneous analysis and identification of all genetic alterations in a single approach with possible turnaround times of 1 week.
Aim: Evaluate the accuracy of WGS and WTS in providing all relevant genetic information in a clinical setting.
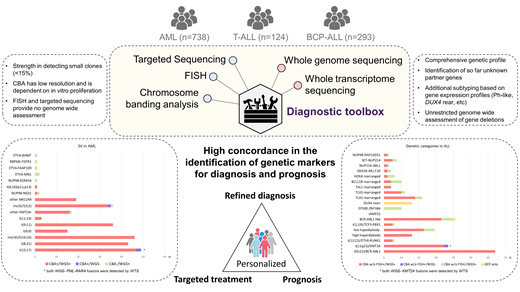
Patients and Methods: The cohort comprised 738 AML, 293 BCP-ALL and 124 T-ALL. The diagnosis was established following WHO guidelines. WGS (100x, 2x151bp) and WTS (50 Mio reads, 2x101bp) were performed on a NovaSeq instrument. Variants were called with Strelka2, Manta and GATK using a tumor w/o normal pipeline, fusions with Arriba, STAR-Fusion and Manta.
Results: The combination of WGS and WTS detected all chromosomal and molecular abnormalities in the AML and ALL cohorts relevant for disease stratification and prognostication as identified by chromosome banding analysis (CBA) and targeted panel sequencing (TPS). A very high concordance between CBA and WGS was revealed for the detection of balanced structural variants (SV) with the added benefit of WGS to also detect cytogenetically cryptic rearrangements (i.e.: ETV6-MN1, NUP98-KDM5A), which all were confirmed either by FISH or RT-PCR. Fusion calling by WTS identified 96% of the WHO subtype defining rearrangements and detected 20 additional fusion transcripts relevant for disease stratification (e.g. EP300-ZNF384, TCF3-HLF) including 9 fusion transcripts that led to prognostic reassignment or could serve as a potential treatment target. Breakpoints of unbalanced SV can occur in repetitive sequences of the genome, hampering the detection by WGS. However, adding copy number alteration (CNA) calls to the analyses allows also reliable identification of unbalanced SV.
WGS outperformed CBA in cases with insufficient in vitro proliferation due to suboptimal pre-analytics (i.e. longer transport time) and identified 36 chromosomal aberrations in 12 cases with CBA not evaluable. WGS's independence of in vitro cell proliferation was most impactful in ALL: 40 T-ALL cases showed a normal karyotype according to CBA. WGS detected SVs in 16 (40%) and CNAs in 20 (50%) of these cases, confirming the normal karyotype for only 9 samples. In the BCP-ALL cohort, CNV analysis identified 29 low hypodiploid and 16 high hyperdiploid karyotypes, 6 of which were missed by CBA. Due to the higher resolution and unrestricted, genome-wide assessment, WGS detected relevant gene deletions (RB1, ERG, PAX5, CDKN2A, IKZF1, ETV6, BTG1) in 59% of ALL cases, providing additional diagnostic and prognostic information. In the AML cohort CBA and WGS detected 795 CNA concordantly. In addition WGS called 54 CNA with size 1-5 MB (below the detection limit of CBA), i.e. 3 BCOR deletions in inv(3)(q21q26) cases and 67 CNA with size > 5 MB, which were missed by CBA. 35 CNA were missed by WGS due to small clone sizes (median 6% as determined by FISH). WGS detected copy neutral loss of heterozygosity (CN-LOH) in AML most frequently on 21q (n=17), 4q (n=15), 13q (n=15), 11q (n=13) and in T-ALL on 9p (n=19), mostly encompassing CDKN2A/B deletions. Expression profiling provided additional diagnostic information for 57 ALL cases (41 BCR-ABL1-like, 16 DUX4 rearranged) that can only insufficiently be obtained by WGS or CBA. WGS reliably detected all gene mutations with a VAF > 15% (n = 647) identified by TPS encompassing especially all mutations in genes relevant for WHO diagnosis and prognostication. 26/171 mutations with a VAF < 15% were missed by WGS. Evaluation of WGS data for 121 genes recurrently mutated in hematologic neoplasms revealed an additional 2 mutations per sample on average (range: 0-9) which might qualify as targets for therapy.
Conclusions: WGS and WTS provide all necessary genetic information to accurately determine the diagnostic and prognostic subgroup according to WHO and ELN guidelines in AML and ALL. Compared to today's gold standards, these novel methods provide a comprehensive genome wide characterization with higher resolution that directly identifies genes of impact, offering the basis for targeted treatment selection and monitoring of residual disease. Both can be implemented with automated analysis pipelines, consequently reducing time and error rates.
Haferlach: MLL Munich Leukemia Laboratory: Other: Part ownership. Kern: MLL Munich Leukemia Laboratory: Other: Part ownership. Haferlach: MLL Munich Leukemia Laboratory: Other: Part ownership.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal